
This field of research aims to define how metabolic pathways and subcellular compartments act in an integrated way, both to regulate plant life and their response to environmental stresses, and to determine the accumulation of proteins and metabolites for food and human health. Part of these investigations are: 1) the identification of genes involved in determining the architecture, trafficking and metabolism of plant cell, also in response to external stimuli; 2) the characterization of the functional and physiological role of the encoded proteins. These studies are carried out through experimental approaches that use in vivo, in vitroand in silico models.


La fotosintesi è il processo mediante il quale la luce solare è convertita in potenziale chimico, utilizzabile dagli organismi, e rappresenta pertantp il sito primario si produzione energetica nella biosfera. In condizioni ottimali le relazioni di foto-conversione procedono con una resa quantica estremamente elevata (80-99%). Lo studio di questi sistemi biologici può quindi rappresentate un paradigma utile per l’implementazione di materiali e strategie che li imitino per applicazioni fotovoltaiche e fotocatalitiche. D’altro canto, in condizioni naturali, a causa di variabilità e fluttuazioni ambientali, la resa fotosintetica è generalmente più bassa di quella ottimale e questo può portare a una riduzione della produttività, soprattutto per quanto riguarda le colture agricole. Pertanto, lo studio dei processi, come la raccolta della luce e il controllo dell’efficienza di raccolta, potrebbe portare ad un miglioramento della produttività delle colture ed essere quindi vantaggioso dal punto di vista sociale.
Il progetto “Enhancing Photosintesi” si propone di affrontare alcuni di questi aspetti, studiando sia i fattori che influenzano la produttività delle colture/piante, sia i loro meccanismi molecolari, sia i meccanismi fondamentali di conversione dei fotoni nei fotosistemi con l’obiettivo di trasferire alcune delle loro caratteristiche in sistemi biomimetici artificiali.
Il progetto prevede inoltre lo sviluppo di una piattaforma spettroscopica (ottica) per lo studio dei processi fotosintetici sia in(super)complessi clorofilla-proteina isolati, sia in sistemi fotosintetici intatti così come in sistemi sintetici artificiali (anche allo stato solido). La strumentazione coprirà oltre ad un ampio intervallo temporale (dai femto ai millisecondi) anche una ambia banda spettrale (dal vicino UV al vicino IR). Questa piattaforma sarà basata sia su strumentazioni già esistenti presso il CNR Milano e al PoliMI, le cui caratteristiche saranno sostanzialmente estese/implementate per renderle adatte allo studio di campioni molto diversificati e complicati da studiare a causa dell’elevata dispersione della luce che presentano. E’ fine del progetto rendere questo “hub” spettroscopico aperto che rappresenti una struttura di duratura che possa aggregare e promuovere la ricerca in fotosintesi, e su argomenti strettamente correlati, a livello regionale e possibilmente nazionale.


Proteins – large molecules central to life – adopt a specific shape to perform their activities. This process is called folding and, for some proteins, it happens in a cell organelle called Endoplasmic Reticulum (ER). In the ER, a wonderfully efficient machinery retains newly made proteins until they are folded properly. This quality control is of great help to healthy people, but in individuals carrying a DNA mutation in a secreted protein gene, the same system causes disease. What happens in these patients is that the quality control system recognises a defect in a mutated protein, which then remains stuck in the ER, even if the defect is small and the patient would still profit from the protein’s exit from the ER, because of its residual activity (“responsive mutant”). Terrible disease ensues. We shall study in the laboratorywhat the quality control does to such responsive mutants, following their trajectories in human cells in which the quality control checkpoint has been deleted. If, in these conditions, the responsive mutant can escape the ER, then a drug that loosens the same checkpoint should have therapeutic potential in a broad range of rare diseases. We selected a panel of ten such defective proteins, hand-picked from the set of 6,000 or so human proteins whose function is little or completely unknown (“Tdark proteins”). This choice has the additional advantage that we shall contribute basic knowledge about Tdarks, towards future treatment of patients whose rare disease is presently not understood let alone cured.
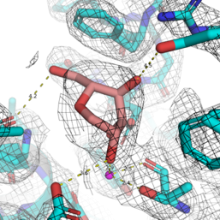
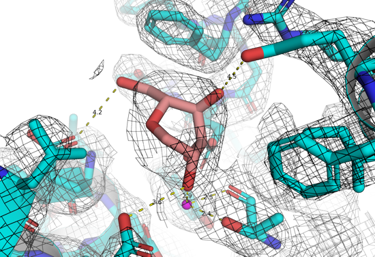
Endoplasmic reticulum-associated degradation (ERAD) is a fundamental strategy of the eukaryotic cell that enables it to degrade all glycoproteins that fail to fold properly – to prevent a build-up of misfolded glycoproteins from clogging the RE. ERAD is initiated by its control enzyme, Endoplasmic Reticulum Degradation Mannosidase (EDEM), which is able to recognise any terminally misfolded glycoprotein, and flag it for degradation. The specific objective of this project is to identify small molecular ligands of HsEDEM3 that bind to the catalytic and/or allosteric sites, and to assay their inhibitory power in the cell. These ligands will constitute starting points for future medicinal chemistry aimed at developing specific inhibitors for HsEDEM3. By Fragment Based Lead Discovery screening, we aim to identify fragments that bind to the catalytic site of HsEDEM3 and compounds that may interfere with the interaction of the catalytic domain with the C-terminal domains of the same protein. The potential long-term outcome of this work would be an allosteric inhibitor or an inhibitor of the catalytic site of HsEDEM3 to be included in antiviral, anti-cancer studies or for the therapy of certain rare diseases. Our aims are: A: Cloning, expression and purification of human EDEM3 GH47 catalytic domain soluble constructs; B: Crystallisation of human EDEM3 GH47 catalytic domain soluble constructs obtained in 1; C: Discovery of ligands of the EDEM3 GH47 catalytic domain soluble constructs (via Fragment-Based Lead Discovery with the crystals from 2); D: Human HsEDEM3 in vitro and cellular inhibition assays to test if the EDEM GH47 ligand fragments obtained in 3. are EDEM3 inhibitors.


In a Drosophila model, the promotion of ER-associated degradation (ERAD) by increasing the levels of ERAD-enhancing α-mannosidase-like proteins (EDEMs) was found to provide protection against chronic ER proteinopathy without causing any toxicity (10.1016/j.devcel.2017.05.019). This protective effect was observed without affecting the UPR gene expression network. Interestingly, as the brain aged, ERAD activity in the fruit fly decreased, but upregulation of EDEMs countered age-related behavioural decline and extended lifespan. Notably, the mannosidase activity of EDEMs was not necessary for these protective effects. Consequently, boosting EDEM function in ERAD holds promise as a potential therapeutic target for chronic diseases. Our recent Cryo-EM structure of an EDEM:PDI complex suggests that the redox activity of the complex is important for its function: clients may be recruited to the complex via mixed disulfides to the PDI. We shall use protein chemistry and structural biology to confirm that the redox activity of the ERAD misfolding checkpoint is important, which in turn may explain the observed mannosidase-independent role of EDEMs in aging.
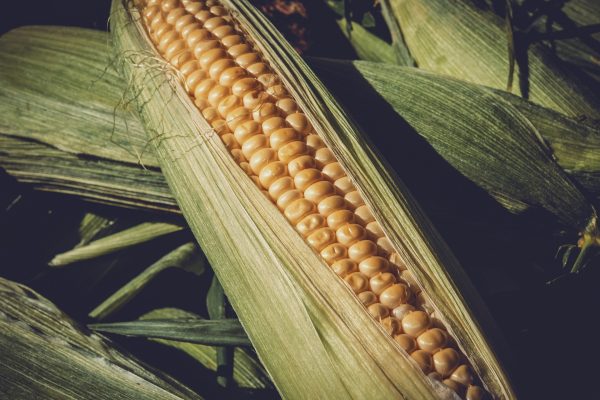


The seed of the maize plant (Zea mays, aka corn) contains zeins, seed storage proteins located within its cells’ Endoplasmic Reticulum (ER) lumen. The maize major seed storage protein, Zea mays 27g-zein (Zm27g-zein), polymerises in the maize seed ER to form large cages – which then accumulate in their interior other maize seed storage proteins, thus providing structural scaffold to the protein body (PB) which is the major food source of the seed. The N-terminus of Zm27g-zein is necessary for the polymerisation to form the PB cage – and the C-terminus of the protein folds as a 2S-albumin fold – but no structures are available for the whole protein nor for either domain in isolation. Most importantly, the way Zm27g-zein gives rise to the molecular structure of the PB cage is completely unknown, including questions such as what intermolecular Zm27g-zein : Zm27g-zein covalent and non-covalent interactions form the PB cage; are there any pores in the PB cage and if so of what size; or the relative orientation of the N- and C-termini in the context for the fully formed PB cage (are Zm27g-zein N-/C- termini in the PB interior, or the exterior, or both?). These questions about the Zm27g-zein PB cage are very important for applications using in the fields of food science, recombinant protein expression, pharmacology and nanotechnology. We shall answer these questions using confocal microscopy, X-ray crystallography and Cryo-EM.






The project includes:




The BIO-ECO project, coordinated by the Department of Biology, Agriculture and Food Sciences of CNR, involves a multi-disciplinary team of researchers belonging to nine Institutes spread all over the country. IBBA activities cover:


Heat and drought stress plants. An important role in resistance to this stress is played by an enzyme called Endoplasmic Reticulum Degradation Enhancing Mannosidase (‘EDEM’). This enzyme exists in a heat-resistant form in a thermophilic fungus originally discovered in a pile of horse manure that fermented in the sun at around 60-65 °C. The first part of the project will test the stress resistance of plants expressing that thermoresistant fungus’ EDEM. Such a transgenic plant could be useful for extracting heavy metals and/or inducing the degradation of organic compounds in contaminated soils (phytoremediation) or in agriculture, to increase agricultural productivity under hostile conditions (heat stress or water shortage during flowering and pollination). The second part of the project aims to obtain a 3D model of EDEM – by X-ray crystallography and/or transmission electron cryomicroscopy. This model will help to understand how EDEM works and to design molecules to modulate its catalytic activity. In addition to the structure, the activity of the enzyme will be characterised in vitro, in cellula and in planta.


EU-IBISBA is a distributed research infrastructure concept that will deliver translational research and innovation services to the research community and industry alike. Its aim is to accelerate the movement of knowledge and early stage research results towards maturity and uptake for further development by industrial R&I. EU-IBISBA’s expertise is within the field of industrial biotechnology and focuses on the integrated development of bioprocesses, using a multidisciplinary approach and advanced technology such as synthetic biology.
PREP-IBISBA intervenes just after the addition of EU-IBISBA on the ESFRI roadmap and as a follow-up to an ongoing H2020 INFRAIA starting community project entitled IBISBA 1.0. The aim of PREP-IBISBA is to create the conditions to launch EU-IBISBA. It will deliver all the conceptual elements necessary to finalize the science and technology case and define the business model, establish a long range financial plan and identify alternative legal frameworks suitable to deliver EU-IBISBA. Overall, the work will lead to the creation of a legal entity, preferentially either an ERIC or another suitable legal instrument.

